In 2007, Jean-Marie Droz of the University of Zurich (working along with Anna Beliakova) wrote a Python program to compute the (hat-version) Heegaard-Floer Knot Homology [math]\displaystyle{ \widehat{\operatorname{HFK}}(K) }[/math] of a knot [math]\displaystyle{ K }[/math] (see arXiv:0803.2379). His program is integrated into KnotTheory`, though to run it, you must have Python as well as the Python library Psyco installed on your system.
(For In[1] see Setup)
| In[1]:=
|
?HFKHat
|
| HFKHat[K][t,m] returns the Poincare polynomial of the Heegaard-Floer Knot Homology (hat version) of the knot K, in the Alexander variable t and the Maslov variable m.
|
|
| In[2]:=
|
HFKHat::about
|
| The Heegaard-Floer Knot Homology program was written by Jean-Marie Droz in 2007 at the University of Zurich, based on methods of Anna Beliakova's arXiv:07050669.
|
|
The Heegaard-Floer Knot Homology is a categorification of the Alexander polynomial. Let us test that for the knot 8_19:
In[3]:=
|
hfk = HFKHat[K = Knot[8, 19]][t, m]
|
Out[3]=
|
2 -3 m 5 2 6 3
m + t + -- + m t + m t
2
t
|
In[4]:=
|
{hfk /. m -> -1, Alexander[K][t]}
|
Out[4]=
|
-3 -2 2 3 -3 -2 2 3
{1 + t - t - t + t , 1 + t - t - t + t }
|
The knot 8_19 is the first knot in the Rolfsen Knot Table whose Heegaard-Floer Knot Homology is not "diagonal". Let us test that. The homology [math]\displaystyle{ \widehat{\operatorname{HFK}}(K) }[/math] is "on diagonal", iff its Poincare polynomial, evaluated at [math]\displaystyle{ m=1/t }[/math], is a monomial:
In[5]:=
|
Select[AllKnots[{3, 8}], (Head[HFKHat[#][t, 1/t]] == Plus) &]
|
Out[5]=
|
{Knot[8, 19]}
|
In[6]:=
|
hfk /. m -> 1/t
|
Out[6]=
|
4 -2
-- + t
3
t
|
The (mirrored) Conway knot K11n34 and the (mirrored) Kinoshita-Terasaka knot K11n42 are a mutant pair, and are notoriously difficult to tell apart. Let us check that an array of standard knot polynomials fails to separate them, yet [math]\displaystyle{ \widehat{\operatorname{HFK}} }[/math] succeeds:
In[7]:=
|
K1 = Knot["K11n34"]; K2 = Knot["K11n42"];
test[invt_] := (invt[K1] =!= invt[K2]);
test /@ {
Alexander, MultivariableAlexander, Jones, HOMFLYPT, Kauffman, Kh, HFKHat
}
|
Out[7]=
|
{False, False, False, False, False, False, True}
|
Indeed,
In[8]:=
|
{HFKHat[K1][t, m], HFKHat[K2][t, m]}
|
Out[8]=
|
2 1 1 3 3 3 3
{3 + - + ----- + ----- + ----- + ----- + ---- + --- + 3 t + 3 m t +
m 4 3 3 3 3 2 2 2 2 m t
m t m t m t m t m t
2 2 2 2 3 3 3
3 m t + 3 m t + m t + m t ,
6 1 1 4 4 2 2 2
7 + - + ----- + ----- + ---- + --- + 4 t + 4 m t + m t + m t }
m 3 2 2 2 2 m t
m t m t m t
|
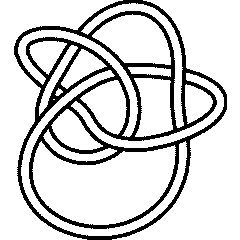
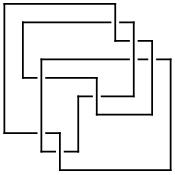
On July 6, 2006, User:AnonMoos asked User:Drorbn if he could identify the knot in the left hand side picture below. At the time it was impossible using the tools available with KnotTheory` - using any of many invariants, the answer can be found to be either the mirror of K11n34 or the mirror of K11n42, but KnotTheory` couldn't tell which one it is (though of course, it is possible to do it "by hand"). The 2007 addition [math]\displaystyle{ \widehat{\operatorname{HFK}} }[/math] does the job, though. Indeed, we first extract the mystery knot's DT (Dowker-Thistlethwaite) Code using the picture on the right hand side below, then compute [math]\displaystyle{ \widehat{\operatorname{HFK}} }[/math], and then search for it within the [math]\displaystyle{ \widehat{\operatorname{HFK}} }[/math]'s of all knots with up to 11 crossings:
In[9]:=
|
K3 = DTCode[6, 8, 14, 12, 4, -18, 2, -20, -22, -10, -16];
|
In[10]:=
|
H = HFKHat[Mirror[K3]][t, m]
|
Out[10]=
|
2 1 1 3 3 3 3
3 + - + ----- + ----- + ----- + ----- + ---- + --- + 3 t + 3 m t +
m 4 3 3 3 3 2 2 2 2 m t
m t m t m t m t m t
2 2 2 2 3 3 3
3 m t + 3 m t + m t + m t
|
In[11]:=
|
Select[AllKnots[], HFKHat[#][t, m] == H &]
|
Out[11]=
|
{Knot[11, NonAlternating, 34]}
|
And so the mystery knot is the Conway knot, the mirror of K11n34.